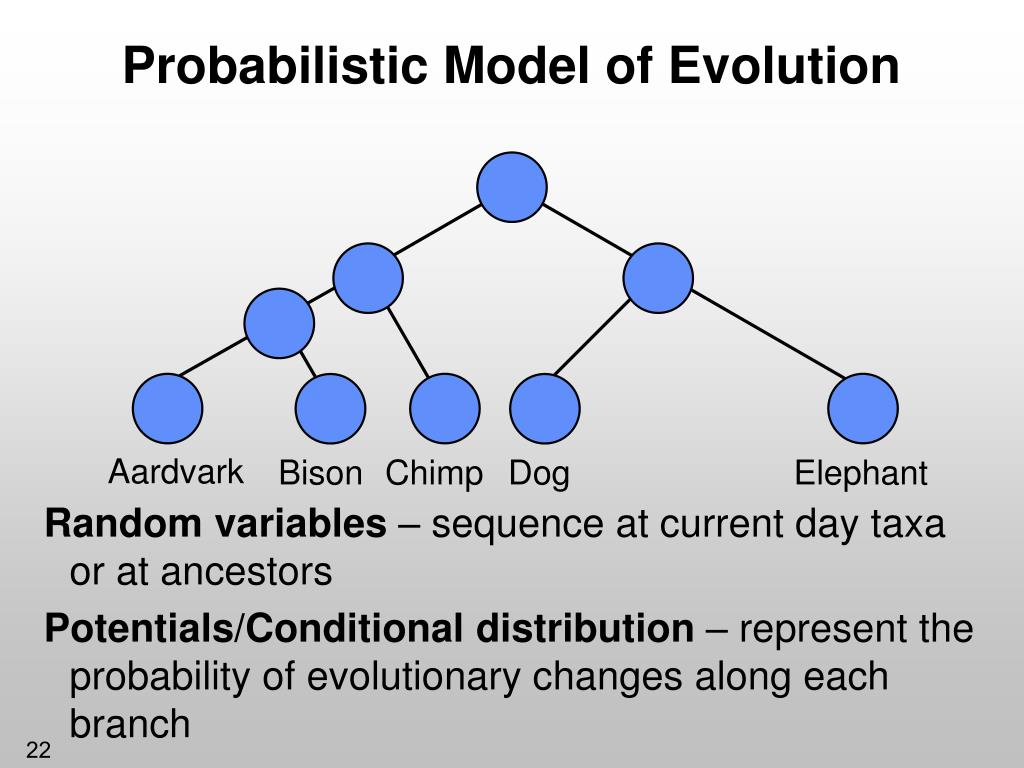
Noticeably, non-homogeneity cannot be simulated by Seq-Gen or PAML, even if these phenomena are all known to affect the evolution of many data sets. However, widely-used simulation programs cannot be easily tuned to precisely reproduce the peculiar evolution of a particular data set. Both for investigating reconstruction methods and for parametric bootstrapping, it is highly desirable that simulation methods model as precisely as possible the conditions that shaped biological sequences through evolution. In this last case, a model has a good fit to a particular data set if the alignments it generates have properties similar to the properties of the real alignment. Similarly, simulations have been used to compare topologies with respect to an alignment, or to assess the fit of a model to a particular data set. For instance, simulations have shown that maximum likelihood methods often more accurately reconstructed the evolution of an alignment than distance or parsimony methods, but could also fail in conditions where compositional biases (a condition here referred to as non-homogeneity) or rate heterogeneity along branches (a phenomenon named heterotachy, ) were too intense. In phylogenetics, simulations have been widely used to study the robustness of inference methods and have been involved in parametric bootstrapping. We furthermore illustrate the use of such general models for parametric bootstrapping, using tests of non-homogeneity applied to an already published ribosomal RNA data set. We show that the general implementation introduced here can accommodate virtually any type of non-homogeneous models of sequence evolution, including heterotachous ones, while being computer efficient. 
These programs allow the user to describe non-homogeneous models through a property file with a simple yet powerful syntax, without any programming required. The first one, Bio++ Maximum Likelihood (BppML), estimates parameters of any non-homogeneous model and the second one, Bio++ Sequence Generator (BppSeqGen), simulates the evolution of sequences from these models. Two programs that use these classes are also presented. It is available as dedicated classes in the Bio++ libraries and can hence be used in any C++ program. We hereby present a general implementation of non-homogeneous models of substitutions. Several programs implementing such models have been developed, but they are limited in their possibilities: only a few particular models are available for likelihood optimization, and data sets cannot be easily generated using the resulting estimated parameters. It has often been observed that homologous sequences can vary widely in their nucleotide or amino-acid compositions, revealing that sequence evolution has changed importantly among lineages, and may therefore be most appropriately approached through non-homogeneous models. Such simulation procedures are needed to estimate the null-distribution of complex statistics, an approach referred to as parametric bootstrapping, and are also used to test the quality of phylogenetic reconstruction programs. Accurately modeling the sequence substitution process is required for the correct estimation of evolutionary parameters, be they phylogenetic relationships, substitution rates or ancestral states it is also crucial to simulate realistic data sets.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
